A kinetic model predicts SpCas9 activity, improves off-target classification, and reveals the physical basis of targeting fidelity
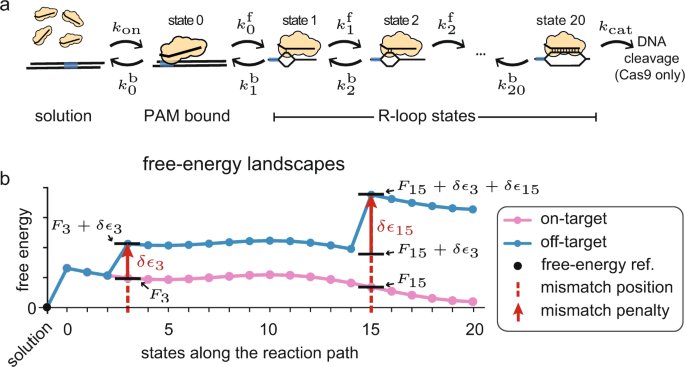

Prediction of off-target effect and gRNA specificity with MOFF a A

A kinetic model improves off-target predictions and reveals the physical basis of SpCas9 fidelity

A kinetic model predicts SpCas9 activity, improves off-target classification, and reveals the physical basis of targeting fidelity

A cleavage rule for selection of increased-fidelity SpCas9 variants with high efficiency and no detectable off-targets

A kinetic model predicts SpCas9 activity, improves off-target classification, and reveals the physical basis of targeting fidelity

Prediction of the sequence-specific cleavage activity of Cas9 variants

A kinetic model predicts SpCas9 activity, improves off-target classification, and reveals the physical basis of targeting fidelity
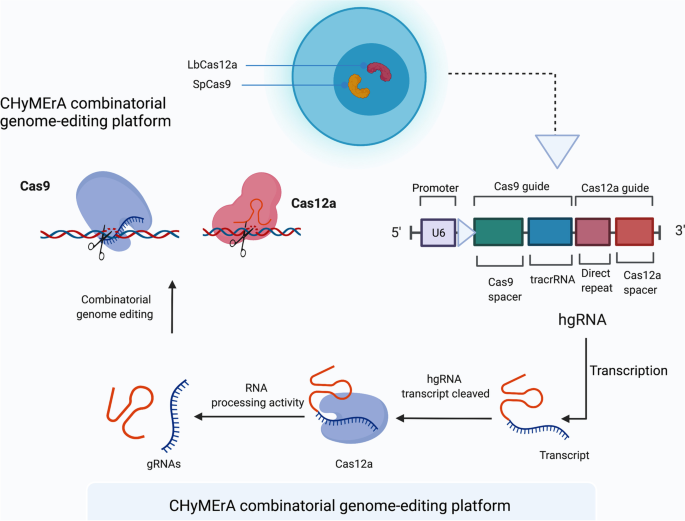
Strategies to overcome the main challenges of the use of CRISPR/Cas9 as a replacement for cancer therapy, Molecular Cancer

A kinetic model improves off-target predictions and reveals the physical basis of SpCas9 fidelity

Strategies to overcome the main challenges of the use of CRISPR/Cas9 as a replacement for cancer therapy, Molecular Cancer

Coordinated Actions of Cas9 HNH and RuvC Nuclease Domains Are Regulated by the Bridge Helix and the Target DNA Sequence

A quantitative model for the dynamics of target recognition and off-target rejection by the CRISPR-Cas Cascade complex

A quantitative model for the dynamics of target recognition and off-target rejection by the CRISPR-Cas Cascade complex

Negative DNA supercoiling induces genome-wide Cas9 off-target activity - ScienceDirect




-980x980.jpg)




